| |
Tandem chimerism as a means to increase protein complexity in the human genome
G. Parra, A. Reymond, N. Dabbouseh, E.T. Dermitzakis, R. Castelo, T.M. Thomson, S.E. Antonarakis, and R. Guigó*.
Genome Research, in press
*To whom correspondence should be adressed.
In this site we describe and provide all the programs and data used to predict chimeric transcripts in the human and mouse genomes.
|
EST whole genome analysis
|
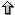
|
First, we searched fusion transcripts in the complete genomes using
expression evidence. In two adjacent non-overlapping coding genes, we
looked for ESTs that alt least partially overlap coding regions in
each of the two adjacent genes. This experiment has been performed in
the human and the mouse genome.
- Human Apr. 2003 Assembly
UCSC Genome Browser
Of the 46 cases in which the ORF was conserved, we tested 32 cases in
which the TIC did not include additional "intergenic" exons. Of these,
11 yielded specific amplification products.
NM1
CODE |
Oligo 1
|
NM2
CODE |
Oligo 2
|
SEQ
CNF |
BR |
HE |
KI |
SP |
LI |
CO |
SI |
MU |
LU |
ST |
TE |
PL |
| NM_007221 |
TCCCAGAGAAGGATCTGCAC |
NM_000711 |
GTGCAGAGTCCAGCAAAGGT |
OK |
1 |
1 |
0 |
1 |
1 |
0 |
0 |
0 |
0 |
1 |
0 |
0 |
| NM_000269 |
GAGACCAACCCTGCAGACTC |
NM_002512 |
CTCGAAGCGCTTGATGATCT |
OK |
1 |
0 |
0 |
0 |
1 |
0 |
0 |
0 |
1 |
1 |
1 |
1 |
| NM_015996 |
CTCCTCCATCGCCATGTT |
NM_003186 |
CACTGCACTATGATCCACTCC |
OK |
1 |
0 |
0 |
0 |
1 |
0 |
0 |
0 |
0 |
0 |
1 |
0 |
| NM_020892 |
GTTCACTGCCAGAGGGTTTC |
NM_030570 |
CGAATGTCAGCCAGTGTCTC |
OK |
0 |
0 |
0 |
0 |
1 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
| NM_002157 |
GGCAGGACAAGCGTTTAGAA |
NM_015387 |
AAAGGATTCATCAGGCCAAT |
OK |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
1 |
0 |
0 |
0 |
0 |
| NM_031925 |
TGCTGGAATGGGAGAAGACT |
NM_016086 |
ATGGAGCTGGTGCAATTGTT |
OK |
1 |
1 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
| NM_153268 |
ATGGAGTTGGTGATGTGCAA |
NM_145753 |
GCTGAGGCTCTCCATCATGT |
OK |
0 |
0 |
0 |
0 |
1 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
| NM_172341 |
TCCTGCCTTTTCTCTGGTTG |
NM_019104 |
TCCAGCAGACACTGCAAGAC |
OK |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
1 |
0 |
| NM_000175 |
GCACCAAGATGATACCCTGTG |
NM_032346 |
AATACACCTGCACGACCAGA |
OK |
0 |
1 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
1 |
0 |
0 |
| NM_003838 |
GCCCTTCAATGTTTGGAAAA |
NM_015978 |
TCAGTTCTTTTTCCTTGATCTGC |
OK |
1 |
1 |
0 |
0 |
0 |
0 |
0 |
1 |
0 |
0 |
1 |
0 |
| NM_025247 |
GCCTTGGATATAGCCATGATT |
NM_000690 |
CCTGACAGATGACCTCTCCA |
OK |
0 |
0 |
0 |
1 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
NM1
CODE |
Oligo 1
|
NM2
CODE |
Oligo 2
|
SEQ
CNF |
BR |
HE |
KI |
SP |
LI |
CO |
SI |
MU |
LU |
ST |
TE |
PL |
|
TABLE COLUMNS DESCRIPTION
Here we describe what does contain each column in the previous table:
- Refseq NM code for the first gene.
- Oligo 1 used for the amplification.
- Refseq NM code for the second gene.
- Oligo 2 used fot the amplification.
- SEQ CNF :
RT-PCR succeeded and the sequence of the fusion gene was confirmed.
- Results on tissues: RT-PCR success on each of the mouse tissues tested in this experiment ("green" color meaning success and "red" color for failure). The tissues were: brain (BR), heart (HE), kidney (KI), spleen (SP), liver (LI), colon (CO), small intestine (SI), muscle (MU), lung (LU), stomach (ST), testis (TE), and placenta (PL).
Positive Genes are those having "OK" in the "SEQ CNF" column.
- Mouse mm4 Assembly
UCSC Genome Browser
- Number of contiguous in the same strand :
7,893
(The file contains the genomic coordinates in gff format and the ids of both RefSeqs)
- Pair of genes with at least one EST crossing the boundaries of the two
known refseqs :
1,670
(The file contains the adjacent RefSeqs ids in the fisrt and second fields and all
the ids of all EST that cross both Refseqs)
- Pair of genes with at least one EST crossing the boundaries and
with at least one overlapping exon :
- Original mapping with BLAT: 208
(format: chromosome EST_id RefSeq1_id number_suported_exons
RefSeq2_id number_suported_exons)
- ESTs re-mapped with Spidey:
135
(format: chromosome EST_id RefSeq1_id number_suported_exons
RefSeq2_id number_suported_exons)
- Spidey aligned pairs after filtering the strand alignment and more than
one EST exon crossing both refseqs:
73
- Pair of genes with at least one EST crossing the boundaries and
with at least one overlapping exon for each gene and a ORF crossing
the junction between the two genes : 19
From the set chimeric transcripts we performed RT-PCR in cDNAs
from different tissues and 8 cases could be confirmed experimentally.
The RT-PCR proves and the amplified tissues.
NM1
CODE |
Oligo 1
|
NM2
CODE |
Oligo 2
|
SEQ
CNF |
BR |
HE |
KI |
TH |
LI |
ST |
MU |
LU |
TE |
SK |
EY |
OV |
| NM_008866 |
GGCCGATCAACAGTGCTAA |
NM_011541 |
TCTTTCAGCAAATCCAATGC |
OK |
0 |
0 |
1 |
0 |
1 |
0 |
0 |
1 |
1 |
1 |
0 |
1 |
| NM_010724 |
GGGTGCCCTCTATCCAGAGT |
NM_011530 |
CCTGAGGCCTCCTTCTCTCT |
OK |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
1 |
0 |
0 |
0 |
0 |
| NM_021419 |
CCCACAGTTTCTGCTCCTTC |
NM_028791 |
GACGATGTTCCCTCCACAAG |
OK |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
1 |
0 |
0 |
0 |
0 |
| NM_026380 |
CTCTCCACGGAAGAAGCAAC |
NM_011267 |
CAGGTTCTCCTCGCTGAACT |
OK |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
1 |
0 |
0 |
0 |
0 |
| NM_027903 |
GGTGGTGAATGGAGAGAGGA |
NM_007527 |
GTAGCAAAAAGGCCCCTGTC |
no |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
| NM_028873 |
AGGCCCAACAGTTCAGTACC |
NM_025364 |
TGGTCTCTAAACCACGAGCA |
OK |
0 |
0 |
0 |
1 |
1 |
0 |
0 |
1 |
1 |
0 |
0 |
1 |
| NM_029090 |
GGAGATATTCTAGCCTCCAGCTT |
NM_029738 |
TTCACAAGCCACAGAAGCAC |
OK |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
1 |
0 |
0 |
0 |
0 |
| NM_134247 |
CCACGCTCTTCCTGCCAC |
NM_134246 |
GCAGGTAGGTCACAGCTTCC |
no |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
| NM_138604 |
TGGGCCTACCGTCATTTAAG |
NM_138606 |
GCCACGACTTGGGTAAAGAA |
OK |
1 |
0 |
0 |
1 |
0 |
1 |
0 |
1 |
0 |
0 |
0 |
0 |
| NM_144894 |
CTTTGAGCCTGCAGCTCTTC |
NM_023684 |
TCCACTGGTACCAAACTGTCC |
OK |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
1 |
0 |
0 |
0 |
0 |
| NM_178017 |
CCTAGCCTCCCTCCTGTCTG |
NM_011622 |
CTCCAGTTGCTGGGAGAGTC |
no |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
NM1
CODE |
Oligo 1
|
NM2
CODE |
Oligo 2
|
SEQ
CNF |
BR |
HE |
KI |
TH |
LI |
ST |
MU |
LU |
TE |
SK |
EY |
OV |
|
TABLE COLUMNS DESCRIPTION
Here we describe what does contain each column in the previous table:
- Refseq NM code for the first gene.
- Oligo 1 used for the amplification.
- Refseq NM code for the second gene.
- Oligo 2 used fot the amplification.
- SEQ CNF :
RT-PCR succeeded and the sequence of the fusion gene was confirmed.
- Results on tissues: RT-PCR success on each of the mouse tissues tested in this experiment ("green" color meaning success and "red" color for failure). The tissues were: brain (BR), heart (HE), kidney (KI), thymus (TH), liver (LI), stomach (ST), muscle (MU), lung (LU), testis (TE), skin (SK), eye (EY) and ovary (OV).
Positive Genes are those having "OK" in the "SEQ CNF" column.
|
geneid predictions in the ENODE regions
|
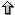
|
Observed cases in the ENCODE regions:
| ENCODE region |
chr |
gene 1 |
transcripts 1 |
gene 2 |
transcripts 2 |
Expressed evidence |
| ENm009 |
chr 11 |
TRIM6 |
NM_001003818
NM_05866 |
TRIM34 |
NM_021616
NM_130389 |
AB039903 |
| ENr233 |
chr 15 |
serf2 |
NM_005770 |
HYpk |
NM_016400 |
AK000438 |
| ENm005 |
chr 21 |
CRYZL1 |
BC033023 |
DONSON |
NM_017613
NM_145794
NM_145795 |
AL157441 |
| ENr223 |
chr 6 |
C6orf148 |
NM_030568 |
AC019205.8-001 |
AK090984 |
BM544101 |
|
The last approach was the prediction of putative new fusion genes
using the ab initio gene preidciton program geneid in the ENCODE
regions. You can find a more detailed information about the selection
and features of the ENCODE regions in the main ENCODE webpage or in the GENCODE webpage. All the
sequences containing pairs of adjacent genes in the ENCODE regions
regions were obtained. geneid
was run on those regions and only predictions that overlapped coding
regions from both genes were considered.
The following subset of sequences have been generated for the experiment:
- Number of initial transcripts (RefSeq, VEGA annotations and Known genes from
UCSC genome browser):
1,288 transcripts.
(This file contains the information of every transcript in
UCSC format )
- Non redundant set of translatable transcripts:
594 transcripts.
(Identificators and corresponding ENCODE region from the filtered previous
set of genes)
- Cluster of overlapping transcripts:
321 clusters.
(The file contains the ENCODE id plus the cluster number GCL_XXX, the
begining and the end of the overlapping transcripts and the ids of the
corresponding ovelapped transcripts)
- Pair of adjacent transcipts (same strand): 165 pairs of clusters.
- geneid predictions overlapping both genes:
126.
(The file contains the ids of the clusters where geneid has predict a protein
that overlaps coding regions of both set of transcripts)
From the previous 126 predictions, 96 where tested by experimental
RT-PCR. The exons corresponding only to the chimeric predictions have
been tested. Three of the chimeric pair of exons predicted by geneid
amplified:
| ENCODE region |
chr |
transcript 1 |
oligo 1 |
transcript 2 |
oligo 2 |
Pred CDS |
Ampli Seq |
| ENm005 |
chr 21 |
NM_144659 |
CCAAGGAGCTGAGAAGAACG |
NM_021254 |
ATACGCTGGCCACAAGAATC |
fasta |
fasta |
| ENm013 |
chr 7 |
AK124057 |
CGGAGAACTTTGTCGGAGAG |
NM_000629 |
TGGAAAAACTAGGGGAAGGA |
fasta |
fasta |
| ENr331 |
chr 2 |
AK023854 |
TGCCTACTTCACTGTCACCA |
BC061909 |
CCAGAGAGGATTGTGCACCC |
fasta |
fasta |
|
TABLE COLUMNS DESCRIPTION
Here we describe what does contain each column in the previous table:
- ENCODE region code.
- Human chromosome where the ENCODE region is.
- Gene 1 code for the first gene.
- Oligo 1 used fot the amplification.
- Gene 2 code for the second gene.
- Oligo 2 used fot the amplification.
- Pred CDS corresponds to the predicted chimeric CDS by geneid.
- Ampli Seq corresponds to the amplified DNA.
|
|